Problem:
Blair Hedges and colleagues have attempted to assign dates to different prokaryotic
metabolisms, among them, oxygenic photosynthesis, with the assumption that
they have adequately accounted for lineage specific rate differences. Their
time constraints for the bacteria are based entirely upon cyanobacterial fossils.
Their data set consists of 54 bacteria and 14 archaea. However, they use only
one cyanobacterium, Synechocystis PCC 6803 in their study. Given what is now
known about the dynamics of cyano genomes, the 16 cyanobacterial genomes currently
available offer an excellent opportunity to test the validity of molecular
clock estimates.
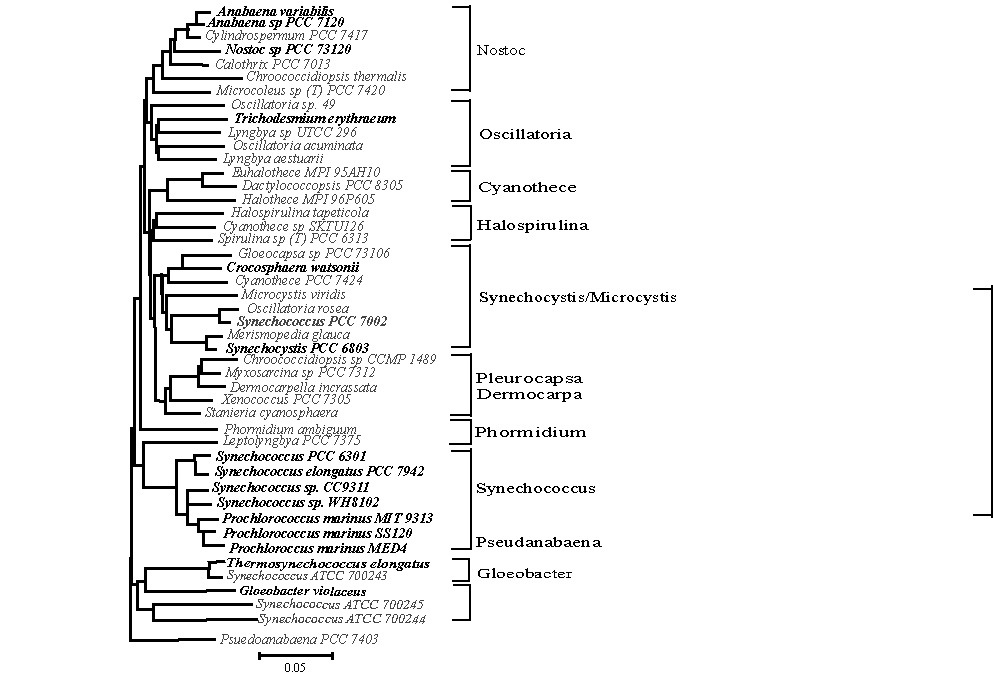
The 16 Cyanobacterial genomes that have been sequenced and are available
span the phylogenetic breadth of the lineage and are depicted in the 16s rRNA
tree.
State of Phylogeny:
The branching order among the cyano groups is ambiguous (Turner et al, 1999).
The nomenclature for the tree follows Turner (other nomenclatures schemes
are elaborated at http://tolweb.org/tree?group=Cyanobacteria&contgroup=Eubacteria).
Biology:
There is evidence that the most deeply branching sequenced genome (Gloeobacter)
is a primitive cyano due to its recurrent position at the base of multiple
gene tress and it 'appears primitive' [it is lacking in photosynthetic membranes
(Nakamura et al, 2003) and PSI is markedly different (Mimuro et al)].
The marine synechococcus group (Syn CC9311 to Prochlo MED4 in tree) has been studied extensively (Rocap, Dufresne, Valout, others.) and the Prochlorococcus are the only free-living genomes reported to be undergoing active genome reduction (Dufresne et al) . This has results in smaller genomes, smaller cell size and genome wide AT skew.
The Nostoc group are a specialized group of cyano. They are multicellular, ie they are filamentous and can differentiate cells into hormogonia (motile cells), akinetes (resting cells) and heterocysts, cells that fix nitrogen. These traits are monophyletic and represent a unique strategy to overcome the problem of the poisoning of nitrogenase by the production of oxygen through photosynthesis (the specialized cells no longer do oxygenic photosynethesis.)
Fossil Record, present state:
There are ambiguous cyanobacteria fossil records for cyano, dating back to
~3.5 Ga (Schopf with major contentions from Brasier that they are not even
biogenic). Stromatolites are argued to be present from very early, and may
or may not be cyano - prior to 2.0, could be anoxygenic photosynthesizers.)
The akinetes from 2.1 appear to be the most robust morphological cyano fossil
(Golubic et al 1995). Brock et al have hopane markers for cyano at 2.7 Ga.

References
Brocks,J.J., Logan,G.A., Buick,R. & Summons,R.E. Archean
molecular fossils and the early rise of eukaryotes. Science 285, 1033-1036
(1999).
Turner,S., Pryer,K.M., Miao,V.P. & Palmer,J.D. Investigating deep phylogenetic
relationships among cyanobacteria and plastids by small subunit rRNA sequence
analysis. J. Eukaryot. Microbiol. 46, 327-338 (1999).
Rocap,G. et al. Genome divergence in two Prochlorococcus ecotypes reflects
oceanic niche differentiation. Nature .424, 1042-1047 (2003).
Brocks,J.J., Buick,R., Summons,R.E. & Logan,G.A. A reconstruction of Archean
biological diversity based on molecular fossils from the 2.78 to 2.45 billion-year-old
Mount Bruce Supergroup, hamersley Basin, Western Australia. Geochimica et
Cosmochimica Acta 67, 4321-4335 (2003).
Dufresne,A. et al. Genome sequence of the cyanobacterium Prochlorococcus marinus
SS120, a nearly minimal oxyphototrophic genome. Proc. Natl. Acad. Sci. U.
S. A 100, 10020-10025 (2003).
Dufresne,A., Garczarek,L. & Partensky,F. Accelerated evolution associated
with genome reduction in a free-living prokaryote. Genome Biol. 6, R14 (2005).
Golubic,S., Sergeev,V.N. & Knoll,A.H. Mesoproterozoic Archaeoellipsoides;
akinetes of heterocystous cyanobacteria. Lethaia 28, 285-298 (1995).
Nakamura,Y. et al. Complete genome structure of Gloeobacter violaceus PCC
7421, a cyanobacterium that lacks thylakoids. DNA Res 10, 137-145 (2003).
Mimuro,M. et al. The secondary electron acceptor of photosystem I in Gloeobacter
violaceus PCC 7421 is menaquinone-4 that is synthesized by a unique but unknown
pathway. FEBS Lett. 579, 3493-3496 (2005).
Battistuzzi,F.U., Feijao,A. & Hedges,S.B. A genomic timescale of prokaryote
evolution: insights into the origin of methanogenesis, phototrophy, and the
colonization of land. BMC. Evol. Biol 4, 44 (2004).