


Next: ims.25
Up: Institute of Mathematical Statistics
Previous: ims.23
IMS
Session Slot: 2:00- 3:50 Monday
Estimated Audience Size: 125-175
AudioVisual Request: None or Two Overheads
Session Title: Information Theory
Theme Session: No
Applied Session: No
Session Organizer: Yu, Bin
Univ of California at Berkeley and
Bell Labs, Lucent Technologies
Address: Statistics Department, 367 Evans Hall,
#3860, University of California,
Berkeley, CA 94720-3860
Phone: (510) 642-2021
Fax: (510) 642-7892
Email: binyu@stat.berkeley.edu
Session Timing: 110 minutes total (Sorry about format):
Opening Remarks by Chair - 5 or 0 minutes
First Speaker - 30 minutes (or 25)
Second Speaker - 30 minutes
Third Speaker - 30 minutes
Discussant - 10 minutes (or none)
Floor Discusion - 10 minutes (or 5 or 15)
Session Chair: Hansen, Mark
Bell Labs, Lucent Technologies
Address:
Phone:
Fax:
Email: cocteau@research.bell-labs.com
1. Information Theoretic Methods in Statistics
Csiszár, Imre,
Mathematics Institue, Hungary Academy of Science
Address: Math. Inst. Hungar. Acad. Sci
H1364 Budapest P.O.Box 127, Hungary
Phone: 011 (36-1) 117-7175
Fax: 011 (36-1) 177-7166
Email: csi@math-inst.hu
Abstract:
While Information Theory uses to a large extent the concepts and
methods of Probability and Statistics, also the other way round,
concepts and methods developed in Information Theory have been
fruitfully applied to various problems of Probability and Statis-
tics. In this talk some such applications will be reviewed, ranging
from Hajek's (1957) information theoretic proof of the dichotomy
theorem for Gaussian measures to Marton's (1997) results on measure
concentration via her information-theoretic method. Topics covered
will include large deviations theory, hypothesis testing, non-para-
metric estimation. The two major inference principles motivated by
Information Theory, viz. Maximum Entropy /or Minimum Discrimination
Information/ and Minimum Description Length would require more time
to discuss at any depth. These will be but briefly illustrated via
relevant examples.
2. Model Selection and Hypothesis Testing by the MDL Principle
Rissanen, Jorma,
IBM Research Division
Address: IBM Research Division,
ARC DPE-B2/802, San Jose, CA 95120-6099
Phone: (408) 927-1813
Fax: (408) 927-2100
Email: rissanen@almaden.ibm.com
Abstract:
The central idea of the MDL (Minimum Description Length)
principle, both in model selection and hypothesis testing, is to
represent a class of models (hypotheses) by a
a universal model capable of imitating the behavior of any model in the
class. The principle then calls for a model class whose representative
assigns the largest probability or density to the observed data. Two
examples of universal models for parametric classes 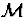 are the
normalized maximum likelihood function
are the
normalized maximum likelihood function
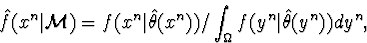
where 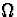 is a small but
simple-to-describe set making the integral finite, and a mixture
is a small but
simple-to-describe set making the integral finite, and a mixture
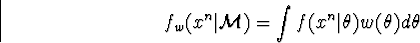
as a convex linear functional of the models.
A Bayes factor 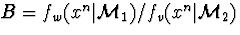 in this
interpretation is the
ratio of mixture representatives of two model classes, and the
Bayesian model comparison reduces to a particular case of the MDL
principle. However, mixtures need not be the best
representatives.
in this
interpretation is the
ratio of mixture representatives of two model classes, and the
Bayesian model comparison reduces to a particular case of the MDL
principle. However, mixtures need not be the best
representatives.
In hypothesis testing, such as in a test of a gaussian singelton class
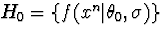 against the remaining class
against the remaining class 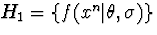 ,we may look for a representative of H1 such that the ratio of its
power to the worst case power of the members in H1 differs least from
unity. The goodness of how well the two hypotheses are separated may
then be assessed by the power of such a minmax representative.
We show that the normalized maximum likelihood is a minmax
representative, while no mixture, taken with respect
to
,we may look for a representative of H1 such that the ratio of its
power to the worst case power of the members in H1 differs least from
unity. The goodness of how well the two hypotheses are separated may
then be assessed by the power of such a minmax representative.
We show that the normalized maximum likelihood is a minmax
representative, while no mixture, taken with respect
to 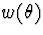 which is symmetric about
which is symmetric about 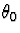 and nonincreasing in
and nonincreasing in
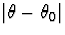 exists with the same minmax property.
exists with the same minmax property.
Time permitting we also discuss other applications of the MDL
principle to model selection.
3. Information and the Clone Mapping of Chromosomes
Yu, Bin,
Univ of California at Berkeley and
Bell Labs, Lucent Technologies
Address: Statistics Department, 367 Evans Hall, #3860,
University of California,
Berkeley, CA 94720-3860
Phone: 510-642-2021
Fax: 510-642-7892
Email: binyu@stat.berkeley.edu
Speed, Terry,
Univ of California at Berkeley
Abstract:
The first step in the Human Genome Project is to assemble DNA fragments,
called clones, to form clone maps, which allow the detailed study of
chromosomal regions of biological interest.
In this talk, we answer biologist Lehrach's question
about how much information is needed to complete a clone map
by formulating a number of different notions of a clone map.
The entropy of each notion (or configuration variable) is tightly
bounded. Our results are useful for planning future mapping efforts.
In particular, it follows that the cosmid
clone mapping for the roundworm requires about 40 times as much
information as that for the bacterium E. coli, and that such
mapping for humans requires about 1,500 times as much information as
that for the bacterium E. coli.
List
of speakers who are nonmembers:



Next: ims.25
Up: Institute of Mathematical Statistics
Previous: ims.23
David Scott
6/1/1998
![]()
![]()
![]() against the remaining class
against the remaining class ![]() ,we may look for a representative of H1 such that the ratio of its
power to the worst case power of the members in H1 differs least from
unity. The goodness of how well the two hypotheses are separated may
then be assessed by the power of such a minmax representative.
We show that the normalized maximum likelihood is a minmax
representative, while no mixture, taken with respect
to
,we may look for a representative of H1 such that the ratio of its
power to the worst case power of the members in H1 differs least from
unity. The goodness of how well the two hypotheses are separated may
then be assessed by the power of such a minmax representative.
We show that the normalized maximum likelihood is a minmax
representative, while no mixture, taken with respect
to ![]() which is symmetric about
which is symmetric about ![]() and nonincreasing in
and nonincreasing in
![]() exists with the same minmax property.
exists with the same minmax property.